RNAcop: Help
Input format
You can either enter a single sequence and a single structure constraint into the corresponding textfields or upload a file which contains multiple sequence-constraint entries. Have a look at the input example and example file from our Submit page.
Sequence
A sequence must be entered as a single line and must only contain the following four canonical bases 'A', 'C', 'G' and 'U'. Optionally, a FASTA-like header tag can be specified as the first line that starts with the character '>', e.g.:Constraint
A constraint must be entered as a single line. You can either specify a constraint for each base which includes the 'no constraint' constraint '.' or you can specify the base where the constraint starts. Bases upstream of the constraint starting position and bases downstream of the last constraint base will automatically be treated as not constrained. A constraint must only contain following characters:
| . | no constraint |
| x | not allowed to pair |
| ( | open base pair |
| ) | close base pair |
| | | must pair with any base |
| < | must pair with any base upstream |
| > | must pair with any base downstream |
This constraint specifies constraints for each base of the sequence, thus the constraint starting position would be base 1. Since a pair '(',')' defines a specific base pair for each opening bracket '(' a closing bracket ')' must be defined. Base pairs which are separated by less than three bases are replaced by no constraint '.'. In addition, non-canonical base pairs, i.e. base pairs other than AU, GC, GU will be removed and replaced by no constraint '.'. If a base pair was replaced a warning message will be printed on the Results page. Instead of adding 215 times '.' in front of the first constraint, you can change Constraint starting position, base to 216.
Sequence-constraint file
A sequence-constraint file can contain multiple entries. Each entry must consist of a unique FASTA-like header starting with character '>' followed by a sequence and then by a constraint of the same length, each given as a single line. When a sequence structure file is used, no constraint starting position can be specified. Therefore, each base must be assigned a constraint (or no constraint '.'), e.g.:
Options
Min/max sizes
You can specify a window of allowed flanking region sizes for suggested flanking regions. Max. 5′ flank size refers to the allowed maximum and Min. 5′ flank size to the allowed minimum size of the flanking region in 5′ direction. Similarly, Max. 3′ flank size refers to the allowed maximum and Min. 3′ flank size to the allowed minimum size of the flanking region in 3′ direction. Please note, for the optimal extension the specified minimum sizes will be ignored since the corresponding log10 probability Pmax should serve as a reference value, but for suggested sizes both minimum and maximum will be considered.
Max. log10 fold-change
Max. log10 fold-change influences which flanking region sizes will be suggested. Inside the user specified window, the maximum log10 probability Pw,max will be determined. Potential flanking region choices are then compared to this value and only values that exhibit a fold-change of equal to or less than the specified value will be considered for flanking region suggestions. Please note, that Pw,max can differ from Pmax, because Pw,max considers both user-specified maximum and minimum sizes for flanking regions.
Email options
Optionally, you can specify an email address and a custom job name if you wish to be informed by email when the computation for your job finishes. Otherwise you can use the given job ID to access the results for your job any time later or simply wait until the computation is done. The Results page will be automatically updated every ten seconds until the computation for the job has finished.
Results
Optimal flanking regions
Results are displayed for each entry separately in case you upload a file with multiple sequence-constraint pairs. First, optimal flanking regions are displayed Optimal flanking regions. Here, the optimal flanking region length in 5′ direction, length 5′ and the optimal flanking region length in 3′ direction, length 3′, as well as the corresponding log10 probability, log10 Pmax. This value should serve as a reference value and is ignoring user-specified minimum sizes for flanking regions. Minimum sizes are considered for Suggested flanking regions.
Suggested flanking regions
Suggested flanking regions are based on the heuristic findgoodregion. In constrast to Optimal flanking regions, only flanking region lengths in the user-specified window are considered. Similarly to Optimal flanking regions, the flanking region length in 5′ direction, length 5′ and the flanking region length in 3′ direction, length 3′, as well as the corresponding log10 probability, log10 Ps is displayed. In addition, a tolerance is given. This value indicates how many nucleotides the size can differ in both 5′- and 3′ direction without decreasing the probability more than the specified fold-change compared to the maximum log10 probability inside the user-specified window. In addition, the corresponding sequence, Suggested sequence, and the according structure constraint Suggested constraint is given.
Other suggested flanking regions are summarized as tab-separated file, Alternative suggestions, including the correpsonding log10 probabilities, radius and area correpsonding to the tolerance, as well as the corresponding sequences. In addition, All pairwise extensions of flanking regions and the corresponding log10 probabilities are given as tab-separated file.
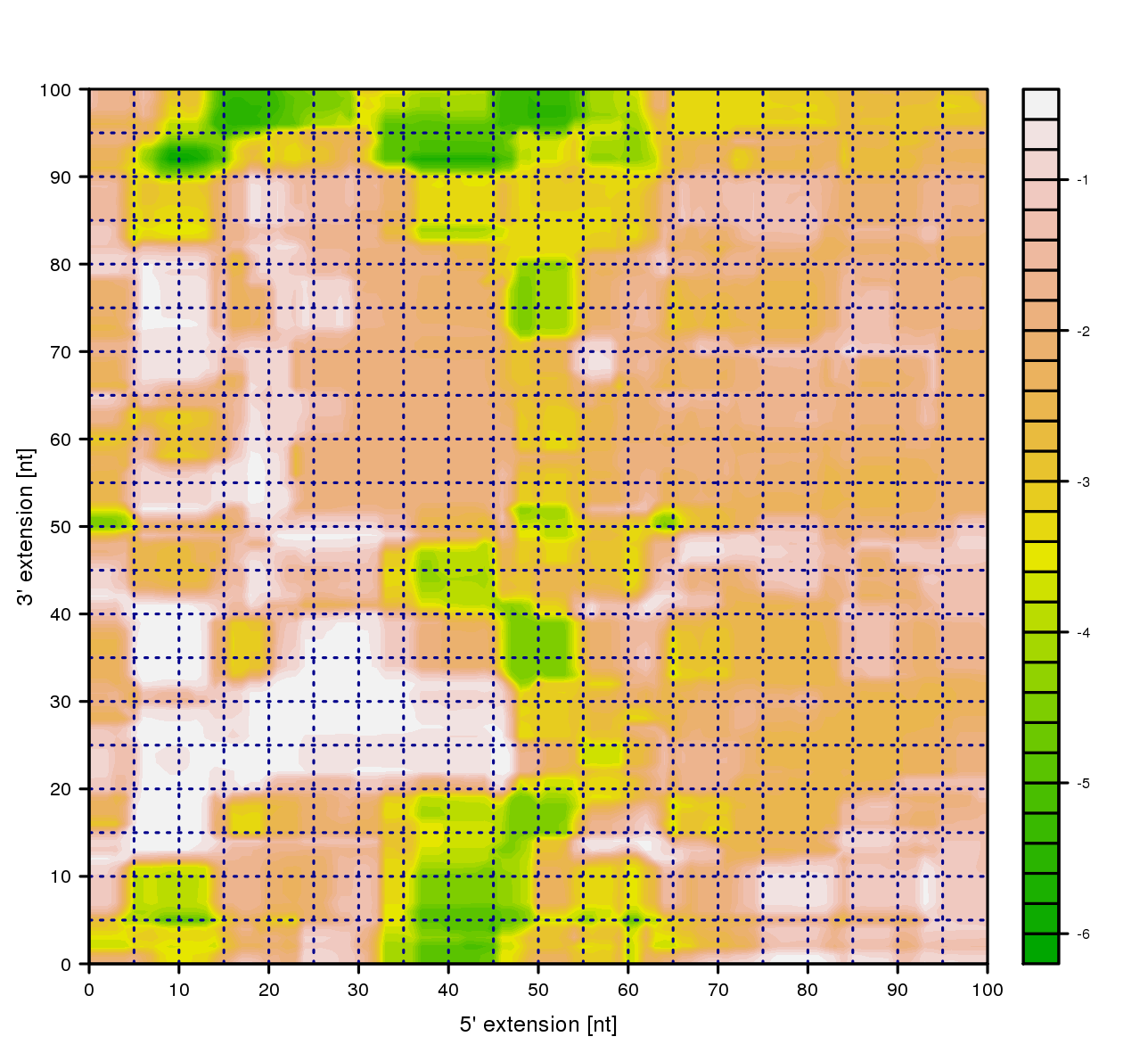
Probability landscape plot
Flanking region extension are visualized as Log10-probability landscape. Where the highest log10-probability is displayed in white and the lowest log10-probability is indicated by a green color. This file can be downloaded as postscript or PNG file.