Center for non-coding RNA in Technology and Health (RTH)
The center aims at developing technologies, computational methods as well as experimental approaches for analysis of the mammalian genome for non-coding RNAs in relation to (inflammatory) diseases. The center will focus on developing these technologies to exploit them and the findings in relation to diabetes. The center consists of a number of national and international partners, with the core located at the Faculty for Health and Medical Sciences of University of Copenhagen .
The people in the center cover a range of expertises including computational biology, RNA bioinformatics, molecular models in diabetes, RNA biology, animal models, functional genomics and high-throughput sequence analysis..
Join us
We are always looking for motivated and talented young scientists as well as projects or colaborations within the areas of the center. Feel free to contact us with suggestions or to ask for more information.
News
CRISPR Cas9 base editing efficiencies
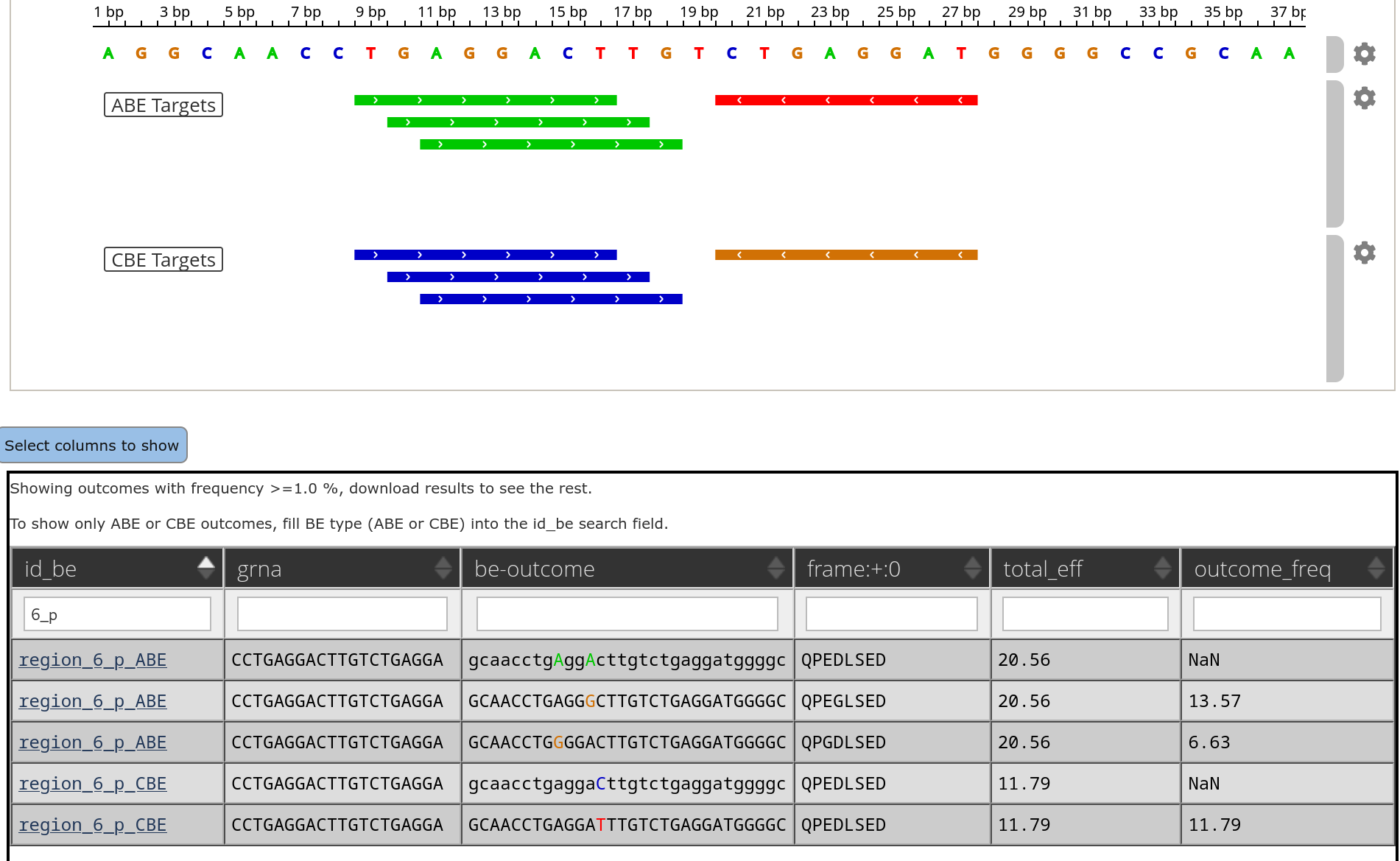
Excited that the collaboration with Yonglun Luo lab on base editing gRNA design has been published in Nature Communications. We generated efficiency and outcome data for ~20000 gRNAs and employed deep learning learning models trained on ours and publicly available data by labeling the individual data sets resulting in a cutting-edge base editor design tool. See the paper here.
Events
2nd CRISPR Genome Engineering and Therapy Symposium
Dec 2nd, 2025 from 10.00 to 17.00
Venue: Copenhagen to be announced
Deadline for registration: Nov 20, 2025
Confirmed speakers:
Bruce Sullenger, University of North CarolinaEric Aird, ETH Zürich
Guillermo Montoya, UCPH
Jacob Nilsson, DCI
Recent resources
CRISPRon-be
Deep learning models simultaneously trained on multiple datasets improve base-editing activity prediction
cyanobacteria CRISPRi
This browsers show the CRISPRi-dCas9 results for a genome-wide knockdown experimental series in Synechocystis sp. PCC 6803 under 4% and 30% CO2 concentrations
Bacillus subtilis PrsA RNAseq data
RNA-seq dataset for a study of the impact of PrsA over-expression on the Bacillus subtilis transcriptome
Research outset
The human genome, made up of DNA, consists of three billion building blocks (nucleotides) where some regions (stretches) are complete genes. We all carry variants of the genes and some cause diseases. Here, the goal is to investigate the specific class of genes, the non-coding RNA genes, in relation to diabetes. The non-coding RNA (ncRNA) genes can be the missing components in diseases that previously have been overlooked.
Our research goal is to develop technologies for ncRNA analysis and to search for functional ncRNAs in relation to diabetes and other (inflammatory) diseases.
Research in details .